Extraction laboratory workflow
Samples that are processed in Biosearch Technologies’ extraction laboratories are barcode-tracked throughout to maintain traceability and ensure sample continuity.
Upon receipt of your samples and required paperwork, Biosearch Technologies will commence laboratory work on your NAP project.
The key stages in the laboratory workflow are:
- Sample checking – Biosearch Technologies checks the condition of all received samples, record any damage and cross-check with the list of expected samples.
- Sample tracking – Biosearch Technologies creates plate layouts for all received samples, ensuring that each individual sample is assigned a specific well location. This enables samples to be fully tracked via barcode as they progress through the laboratory.
- Sample processing – samples are extracted in our service laboratory using standard operating procedures and are eluted according to customer-specific requirements.
- Sample quantification and normalisation – eluted DNA samples are quantified and normalised according to customer-specific requirements.
Sample checking
Received samples are checked upon receipt to ensure that they have been received in good condition. Any damage or contamination is recorded, and the customer is informed. For samples received in tubes, the sample IDs on each tube are cross-checked against the sample manifest that has been provided by the customer to ensure that all expected samples have been received.
Sample tracking
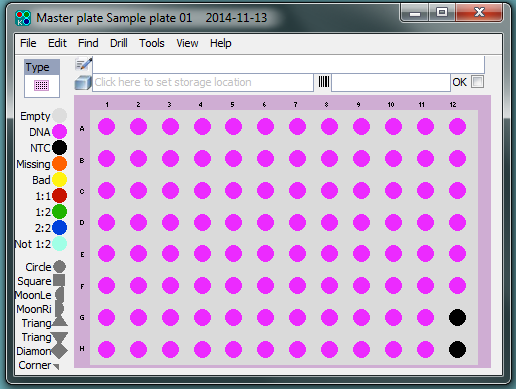

For samples that have been received in plates, a plate map file will have been received from the customer. This plate layout information is stored in Kraken, within the individual project. Plate barcodes are linked to the plate layout information, ensuring that individual samples can be fully tracked as they progress through the laboratory.
For samples that have been received in individual tubes, plate layouts are created within Kraken and each unique sample is assigned a specific well location using the tube barcode. In every 96-well plate layout, two wells are intentionally left empty for controls.
Sample processing
For every NAP project run through our service laboratories, a project manager will be assigned who is responsible for ensuring that the project is processed according to customer requirements and within the required turnaround time. On a project information sheet, the project manager will specify the protocol to be used for extraction, the required buffer volumes, and any customer-specific information regarding sample elution. Laboratory staff will utilise the project information sheet in conjunction with Standard Operating Procedures to perform extraction from the samples.
Extraction in Biosearch Technologies’ service laboratories is performed using our own chemistries. Blood, buffy coat, saliva and plant samples are typically extracted using our sbeadex™ chemistry. Where appropriate, our oKtopure™ robotic platforms are also used.
Sample quantification and normalisation
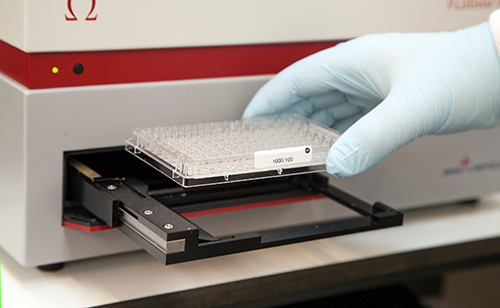
Eluted DNA samples are quantified using UV spectrophotometry (FLUOstar Omega plate reader) as standard. Biosearch Technologies also offer quantification using PicoGreen®, a fluorescent nucleic acid stain that selectively binds to double-stranded DNA. Please view our factsheet and application note that compare DNA quantification values obtained using UV spectrophotometry and PicoGreen analysis.
The increased sensitivity of PicoGreen quantification, compared to UV spectrophotometry, means that it is particularly applicable to quantification of DNA from limited clinical samples with low numbers of cells or to samples from which extraction of DNA is problematic and results in low yield (e.g. some plant seeds/leaf tissue). In addition, PicoGreen quantification can often be a pre-requisite for downstream applications such as microarrays and sequencing. If you wish to include PicoGreen analysis in your DNA extraction project, please advise your sales rep or project manager in advance of commencing your project.
