SNPviewer is a tool that enables genotyping data to be viewed as a cluster plot (see below), but that does not include data analysis or reporting functionality. Genotyping data from a service project run at LGC Biosearch Technologies is typically sent to the customer as a SNPviewer file (in addition to a text file). Opening genotyping service project results in SNPviewer will allow you to view the genotyping clusters plate by plate. Genotyping calls cannot be changed within SNPviewer; if you need to be able to edit the calls then you will require our KlusterCaller software.
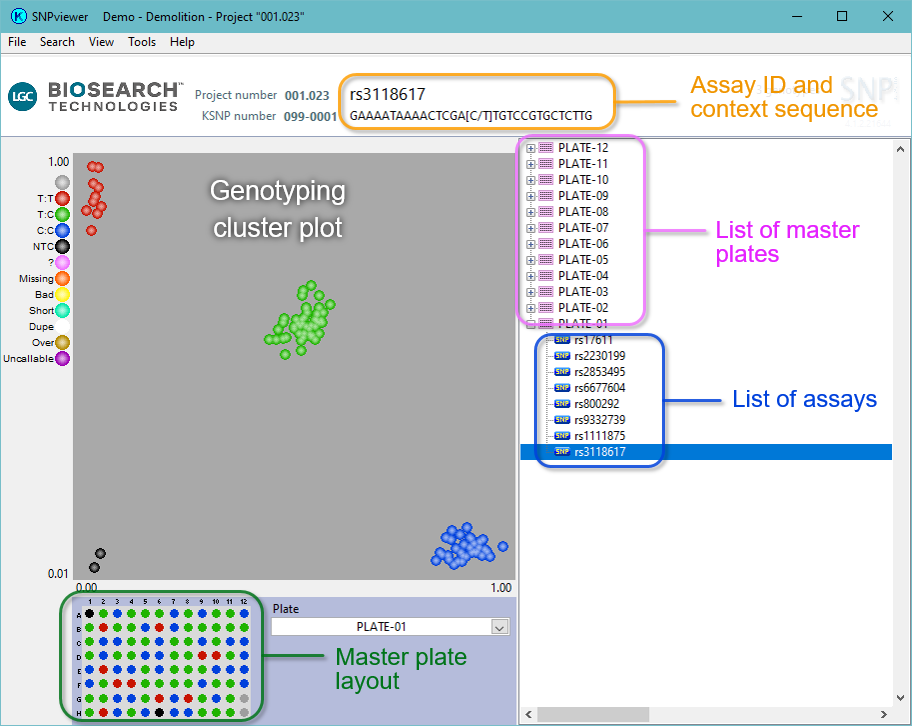
Your genotyping data may be visualised by downloading the SNPviewer PC tool. This will let you graphically view the clusters that group the allele calls. This software will present your results on a treeview by SNP and master plate. The plates may be clicked on to view the data. Calls can not be changed in SNPviewer. If you need to change any calling then you need KlusterCaller.
Minimum PC specification
SNPviewer 64-bits is suitable for computers that are running on Windows 10 64-bits.
Installation instructions
You will need administration rights on your PC to install SNPviewer.
- Click here to download - SNPviewer
- Right click the installer ‘SNPviewer.setup.26546.exe’ and select ‘Run as administrator’. Please note: you should first uninstall any existing SNPviewer-version from your machine. This can be done from ‘Control Panel’ > ‘Programs and Features’. Once you have done this, start the installation as an administrator.
- Work through the installation wizard to install SNPviewer on your machine.